Getting Started
Usage Agreement
Please ensure that you have read Zurada’s System Use Policies and have agreed to it before requesting an account.
HPC system overview
About the cluster
Zurada consists of 119 total nodes out of which, 2 are login nodes, 3 are test nodes used by system administrators, and 112 nodes that can be used for computation. These nodes are distributed in the following queues:
Queue Name |
Number of servers |
Processor |
CPU core frequency |
CPU sockets per node |
CPU cores per socket |
Total CPU cores per node |
Raw memory |
Usable memory |
GPU |
GPU Memory |
GPUs per node |
Local storage per node |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
cpu384g |
74 |
AMD EPYC 9174F |
4.10 GHz |
2 |
16 |
32 |
384 GiB |
376 GiB |
N/A |
N/A |
0 |
480 GB SATA mixed use SSD |
cpu1500g |
6 |
AMD EPYC 9174F |
4.10 GHz |
2 |
16 |
32 |
1.5 TiB |
1.4 TiB |
N/A |
N/A |
0 |
6.4 TB NVMe mixed use SSD |
cpu6000g |
3 |
AMD EPYC 9174F |
4.10 GHz |
2 |
16 |
32 |
6 TiB |
5.8 TiB |
N/A |
N/A |
0 |
6.4 TB NVMe mixed use SSD |
gpu1h100 |
13 |
AMD EPYC 9174F |
4.10 GHz |
2 |
16 |
32 |
1.5 TiB |
1.4 TiB |
NVIDIA H100 NVL |
95830 MiB |
1 |
6.4 TB NVMe mixed use SSD |
gpu2h100 |
10 |
AMD EPYC 9174F |
4.10 GHz |
2 |
16 |
32 |
3 TiB |
2.8 TiB |
NVIDIA H100 NVL |
95830 MiB |
2 |
6.4 TB NVMe mixed use SSD |
hgxh200 |
1 |
Intel Xeon Platinum 8480C |
2.0 GHz |
2 |
56 |
112 |
2 TiB |
1.9 TiB |
NVIDIA H200 NVL |
1128 GiB |
8 |
30 TB NVMe |
cpudev |
3 |
AMD EPYC 9174F |
4.10 GHz |
2 |
16 |
32 |
384 GiB |
376 GiB |
N/A |
N/A |
0 |
480 GB SATA mixed use SSD |
gpudev |
2 |
AMD EPYC 9174F |
4.10 GHz |
2 |
16 |
32 |
1.5 TiB |
1.4 TiB |
NVIDIA H100 NVL |
95830 MiB |
1 |
6.4 TB NVMe mixed use SSD |
The cluster features 200 Gbps NDR Infiniband connectivity between compute nodes.
These nodes are named as follows:
cpusm01,cpusm02, …,cpusm74for cpu-only nodes in queuecpu384g.cpumd01,cpumd02, …,cpumd06for cpu-only nodes in queuecpu1500g.cpulg01,cpulg02,cpulg03for cpu-only nodes in queuecpu6000g.gpusm01,gpusm02, …,gpusm13for gpu nodes in queuegpu1h100.gpumd01,gpumd02, …,gpumd10for gpu nodes in queuegpu2h100.gpulg01for the gpu node in queuehgxh200.
There are also 2 development queues:
cpudevwithcpusm75…cpusm77gpudevwithgpusm14…gpusm15
These development queues have slightly different restrictions, and are available so that users are not waiting for a long job to complete just to run a couple of quick tests.
In order to execute any kind of (scientific) software, such as Ansys, OpenFOAM, GROMACS, or others, on these nodes, users must:
Log into a special node referred to hereinafter as the “login node” (see Section Logging into the cluster for more information).
Submit a job declaration to the scheduling system, Slurm. Slurm ensures fair resource distribution among users by managing node allocation, CPU distribution, memory utilization, and other essential resources based on job requirements.
To prevent interference between users’ jobs, access to nodes is restricted
to users with active jobs running on them. For example, if user “lk01” submits a job and
Slurm allocates cpusm01 for its execution, “lk01” will have exclusive access to log into cpusm01.
About Scientific Software
In Linux, program behavior is influenced by dynamic values called “environmental variables”.
These variables can be created, modified, and removed as needed, shaping the functionality
of programs and services. For example, the PATH variable lists directories where binaries are stored.
When a command is executed, the system searches these directories for the corresponding binary.
If not found, it returns a “command not found” error.
Scientific software like GROMACS or OpenFOAM often define their own environmental variables, which can be complex to manage. To simplify this, the cluster uses Environmental Modules to dynamically adjust users’ environments with modulefiles.
Users can explore available modules with the module available command and load
them using module load modulename.
About Jobs
Users can submit two types of jobs: interactive and batch. Interactive jobs give direct access to the assigned node, allowing users to execute programs manually. Batch jobs run autonomously via shell scripts without user intervention.
Batch jobs remain unaffected by disconnections, while interactive jobs may terminate.
To maintain continuity, users can use a terminal multiplexer like tmux.
Running tmux before starting an interactive job creates
a persistent session that continues even if the connection is lost.
Quickstart
Logging into the cluster
Upon creating an account, users are provided with a username and password, which they can utilize to access the cluster via SSH (Secure Shell Protocol). The procedure entails employing an SSH client from their personal computers to establish a connection with the login node.
Using the command line
Windows (versions 10 and 11) inherently supports an SSH command-line client within PowerShell. Similarly, Mac and Linux based operating systems come equipped with a built-in SSH client accessible via their respective terminals. The basic login process remains consistent across all of these platforms:
Launch the terminal on your personal computer.
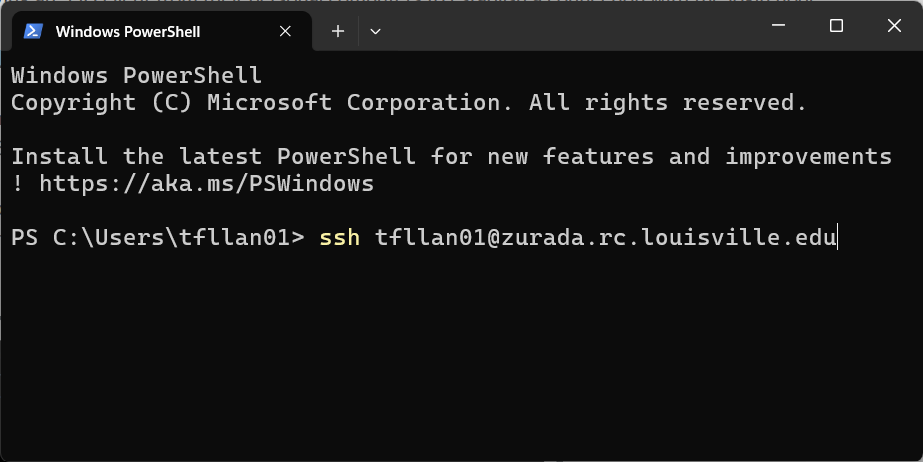
Enter the ssh command using the following format:
ssh username@hostname. In this particular scenario, the hostname is alwayszurada.rc.louisville.edu. For instance, if the user’s name is “lk01”, they would inputssh lk01@zurada.rc.louisville.edu. Here is an example:
Provide your password and press Enter.
Alternatively, users can opt for other popular SSH clients installed on their personal computers, such as MobaXterm and PuTTY. PuTTY boasts a straightforward and user-friendly interface, while MobaXterm offers a tabbed interface with enhanced functionality, including a dedicated file manager that simplifies file management on the cluster and facilitates seamless information transfer between the personal computer and the cluster.
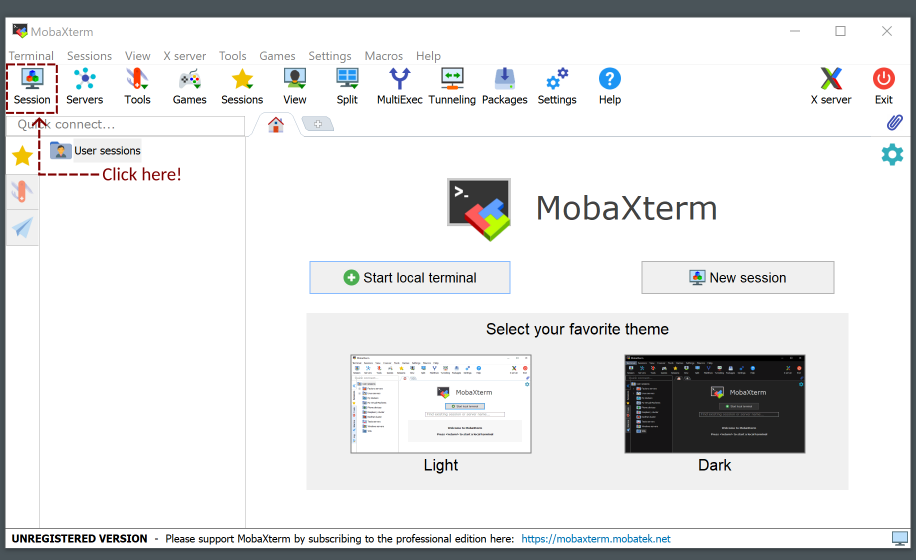
Using MobaXterm
Click on “Session” at the top-left of the window
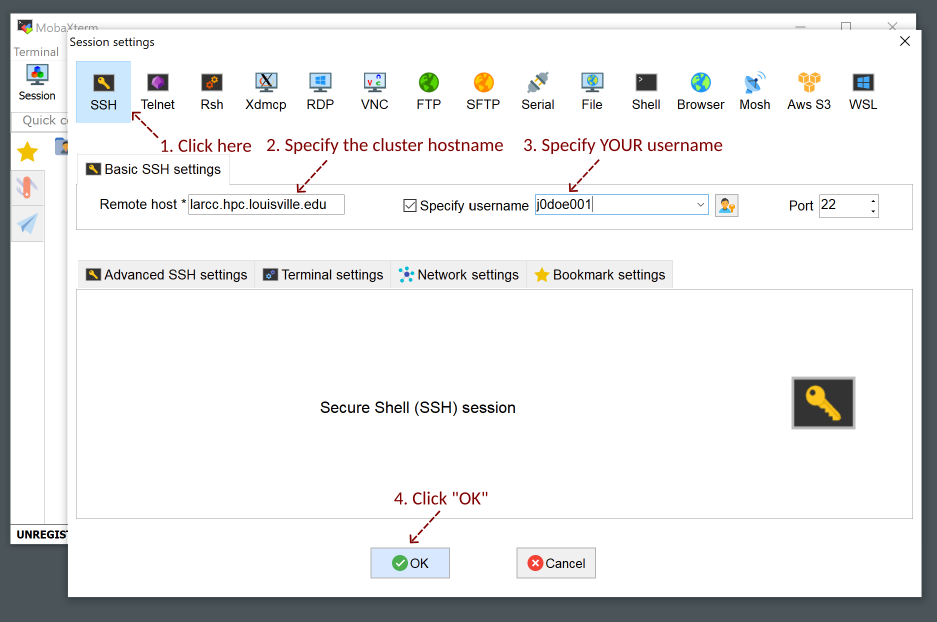
Setup your username and the cluster hostname
zurada.rc.louisville.edu
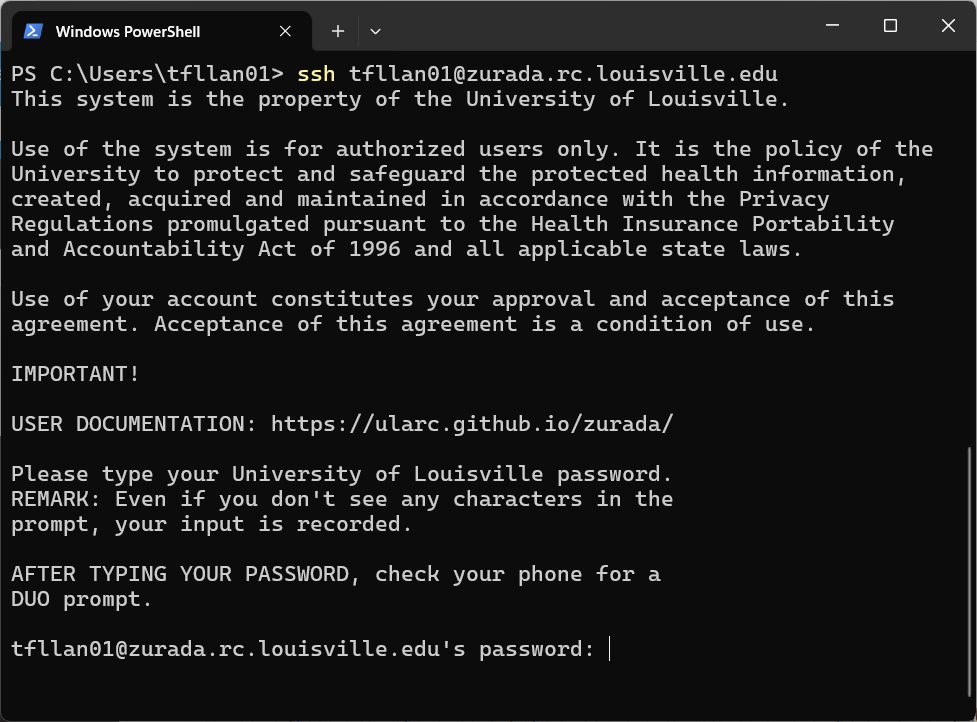
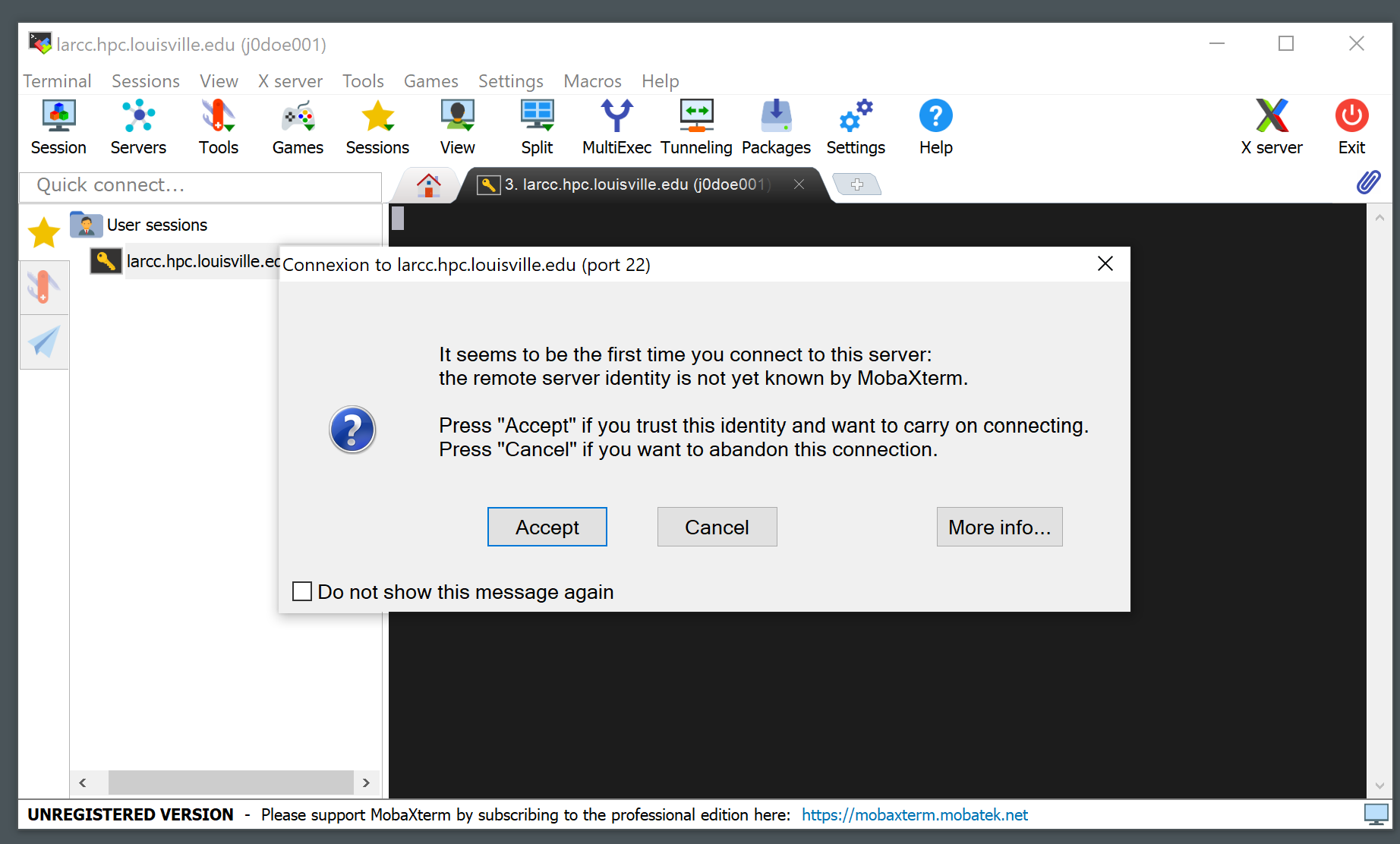
A notice like the one below will appear the first time you connect to the cluster. Click “Accept”.
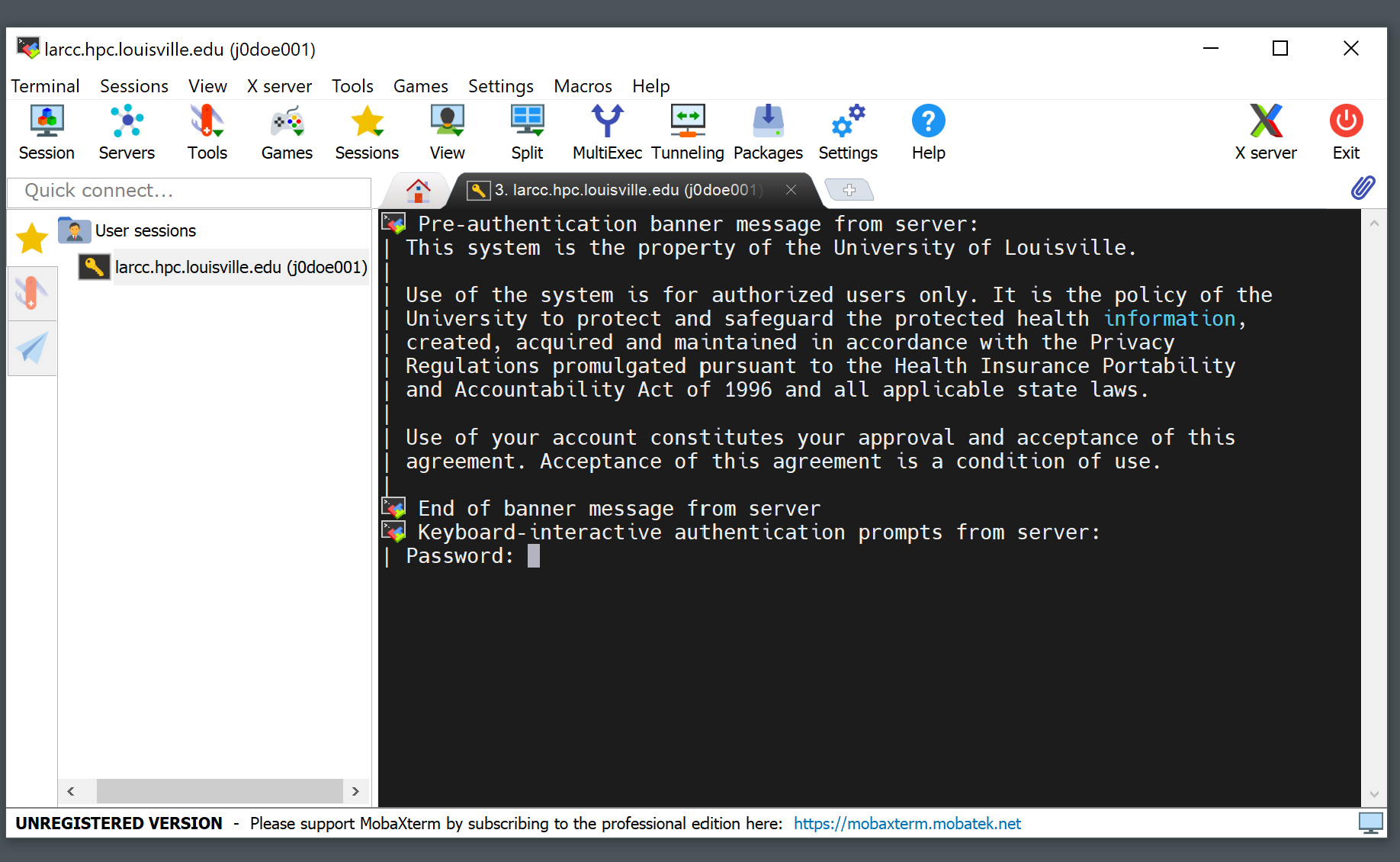
Write your password (it will not be displayed as you type it) and hit Enter
Copying files to/from the cluster
Using the command line
The command scp (available on Windows, Mac and Linux based OSs) is the preferred way
to copy files to and from the cluster. See a comprehensive list of options at the
scp guide. Since a user’s
home directory (/home/<username>, or simply ~) is shared across all nodes, users are encouraged
to use their home directories as a staging area for file transfers.
Example: Assume user “John Doe” is assigned cluster account jd01. The code below
shows how John would copy the file C:\Users\johndoe\Downloads\workload.jou from his
personal computer to his home directory (/home/jd01) in the cluster using the
scp command in Windows PowerShell.
# John could also use ~ instead of /home/jd01. That is, the following is also valid:
# scp C:\Users\johndoe\Downloads\workload.jou jd01@zurada.rc.louisville.edu:~
scp C:\Users\johndoe\Downloads\workload.jou jd01@zurada.rc.louisville.edu:/home/jd01
Suppose John Doe ran a simulation and got the results stored at /home/jd01/results/sim_1_res.dat
in the cluster. If he wants to copy these retults to the folder C:\Users\johndoe\Documents
of his Windows PC, he would execute the command below from a PowerShell session:
# The following is also valid:
# scp jd01@zurada.rc.louisville.edu:~/results/sim_1_res.dat C:\Users\johndoe\Documents
scp jd01@zurada.rc.louisville.edu:/home/jd01/results/sim_1_res.dat C:\Users\johndoe\Documents
Using MobaXterm
Downloading files or folders from the cluster
Locate the “File Explorer” from MobaXterm and navigate towards the location where the file or folder you want to download resides in.
Right click on the file or folder you want to download from the cluster and click on “Download”.
Uploading files or folders to the cluster
Locate the “File Explorer” from MobaXterm and navigate towards the location where you want to upload your files to.
Click on the upload icon within the “File Explorer” and select the file or folder you want to upload.
Using software installed in the cluster
List available software
Use command module avail as shown in the example below:
[user@login01 ~]$ module av
---------------------------------------- /mnt/apps/modulefiles/manual -----------------------------------------
aocc/5.0.0-gcc-11.4.1 miniforge3/25.3.1-gcc-11.4.1
aocc/5.0.0-gcc-12.4.0 (D) mpc/1.3.1-gcc-11.4.1
aocl/5.1.0-aocc-5.0.0 mpfr/4.2.1-gcc-11.4.1
bcftools/1.22-gcc-12.4.0 mvapich2/2.3.7-aocc-5.0.0
binutils/2.45-gcc-11.4.1 nvhpc/25.5
cmake/4.2.1-gcc-12.4.0 openmpi/5.0.5-aocc-5.0.0-gcc-12.4.0
curl/8.16.0-gcc-12.4.0 orca/6_1_0-avx2
gaussian/16-A.03-nvhpc-25.5 r/4.5.1-aocc-5.0.0-gcc-12.4.0
gcc/12.4.0-gcc-11.4.1 slurm-libpmi2/24.11.5-gcc-11.4.1
gmp/6.3.0-gcc-11.4.1 vasp/6.5.1-aocc-5.0.0-aocl-5.1
intel-oneapi/2025.2.1.44 vasp/6.5.1-nvhpc-25.5-mkl-2025.2 (D)
isl/0.27-gcc-11.4.1 xz/5.8.1-gcc-12.4.0
lammps/20240829.2-nvhpc-25.5-aocc-5.0.0 zstd/1.5.7-gcc-11.4.1
matlab/r2025b
Where:
D: Default Module
Load software
Users must load programs with module load <modulename> before launching them.
Multiple programs can be loaded at the same time, but there are cases where two or more may conflict.
For instance, programs openmpi/5.0.5-aocc-5.0.0-gcc-12.4.0 and mvapich2/2.3.7-aocc-5.0.0
cannot be loaded together. An example of this is shown below:
[user@login01 ~]$ module load openmpi/5.0.5-aocc-5.0.0-gcc-12.4.0
[user@login01 ~]$ module load mvapich2/2.3.7-aocc-5.0.0
Lmod has detected the following error: You can only have one mpi module loaded at a time.
You already have openmpi loaded.
To correct the situation, please execute the following command:
$ module swap openmpi mvapich2/2.3.7-aocc-5.0.0
While processing the following module(s):
Module fullname Module Filename
--------------- ---------------
mvapich2/2.3.7-aocc-5.0.0 /mnt/apps/modulefiles/manual/mvapich2/2.3.7-aocc-5.0.0.lua
[user@login01 ~]$
Warning
Programs MUST only be run through slurm, NOT on a login node. Users can test their scripts using an interactive job first and then submit the appropriate batch job (See our Slurm Queueing System Guide for more details).
List currently loaded software
Use command module list as shown in the example below:
[user@login01 ~]$ module load openmpi/5.0.5-aocc-5.0.0-gcc-12.4.0
[user@login01 ~]$ module list
Currently Loaded Modules:
1) gmp/6.3.0-gcc-11.4.1 6) zstd/1.5.7-gcc-11.4.1
2) mpfr/4.2.1-gcc-11.4.1 7) gcc/12.4.0-gcc-11.4.1
3) mpc/1.3.1-gcc-11.4.1 8) aocc/5.0.0-gcc-12.4.0
4) isl/0.27-gcc-11.4.1 9) openmpi/5.0.5-aocc-5.0.0-gcc-12.4.0
5) binutils/2.45-gcc-11.4.1
Note
In addition to openmpi/5.0.5-aocc-5.0.0-gcc-12.4.0, several other programs are listed.
These are dependencies that the module automatically loads alongside OpenMPI.
Dependencies marked with an H are hidden by default.
This means they will not appear when you run the module available command,
even though they are still loaded and available for use.
Unloading software
Use command module unload <modulefile>. This command only unloads the
indicated program, but not its dependencies. To clean the environment and
unload all modules, users should use the command module purge. Example:
[user@login01 ~]$ module load openmpi/5.0.5-aocc-5.0.0-gcc-12.4.0
[user@login01 ~]$ module list
Currently Loaded Modules:
1) gmp/6.3.0-gcc-11.4.1 6) zstd/1.5.7-gcc-11.4.1
2) mpfr/4.2.1-gcc-11.4.1 7) gcc/12.4.0-gcc-11.4.1
3) mpc/1.3.1-gcc-11.4.1 8) aocc/5.0.0-gcc-12.4.0
4) isl/0.27-gcc-11.4.1 9) openmpi/5.0.5-aocc-5.0.0-gcc-12.4.0
5) binutils/2.45-gcc-11.4.1
[user@login01 ~]$ module purge
[user@login01 ~]$ module list
No modules loaded
[user@login01 ~]$
Queues and jobs
The cluster has six main queues named cpu384g, cpu1500g, cpu6000g, gpu1h100, gpu2h100, hgxh200, and two development queues named cpudev and gpudev.
To see information about queues, users can use the
sinfocommand.When users send jobs, they can monitor their job status using the
squeuecommand.To launch an interactive job, users can user the
srun --time=<walltime> --pty /bin/bash -icommand. See Section Starting an interactive job for more information.To submit an unattended job, users can use the command
sbatchas follows:sbatch /path/to/sbatch/script. See Section Submitting batch jobs for more informationTo cancel jobs, users can use the
scancelcommand as follows:scancel jobid
Resource Restrictions
Running applications on the login nodes
Users should avoid running resource-intensive workloads on the login nodes, as this can degrade performance and hinder others from accessing the cluster or submitting jobs. To maintain a stable and fair environment, the Research Computing Team reserves the right to terminate any user processes on the login nodes that are found to negatively impact other users.
Job runtime restrictions
Note
Please note that exceptions to the restrictions described below CAN be made.
If your workload needs to be given more time to run, you need to use more nodes than what is allowed by default, among others, please reach out to us by creating a ticket and we will be happy to evaluate your case.
If the
--timeoption is not specified when submitting a job, a default runtime of 12 hours is imposed on said job. This applies to both interactive and batch jobs.To see runtime restrictions on each partition, please run the
sinfocommand and look at theTIMELIMITcolumn.The number of usable nodes per user in a partition can be seen as follows:
use the
scontrol show partition <partition name>command to print information about the partition. In the output, locate theQoS=line and take note of the value. For instance, in the example below, partitioncpu384guses QoScompute-sm_user_limits.
scontrol show partition cpu384g PartitionName=cpu384g AllowGroups=ALL AllowAccounts=ALL AllowQos=ALL AllocNodes=ALL Default=YES QoS=compute-sm_user_limits DefaultTime=NONE DisableRootJobs=NO ExclusiveUser=NO ExclusiveTopo=NO GraceTime=0 Hidden=NO MaxNodes=8 MaxTime=3-00:00:00 MinNodes=0 LLN=NO MaxCPUsPerNode=UNLIMITED MaxCPUsPerSocket=UNLIMITED Nodes=cpusm[01-77] PriorityJobFactor=1 PriorityTier=1 RootOnly=NO ReqResv=NO OverSubscribe=EXCLUSIVE OverTimeLimit=NONE PreemptMode=OFF State=UP TotalCPUs=2464 TotalNodes=77 SelectTypeParameters=NONE JobDefaults=(null) DefMemPerNode=UNLIMITED MaxMemPerNode=UNLIMITED TRES=cpu=2464,mem=29722000M,node=77,billing=2464
use the
sacctmgr show qos <qos name> format=MaxTRESPUcommand to print the resource restrictions of a qos. For instance, the example below shows that the qoscompute-sm_user_limits, assigned to partitioncpu384g, restricts users to a maximum of 8 nodes at any given time.sacctmgr show qos compute-sm_user_limits format=MaxTRESPU MaxTRESPU ------------- node=8
Users can submit a maximum of 50 jobs across all partitions.
Example: Consider user jd01 submits 2 jobs named A and B such that each job requests 4 nodes from the
cpu384gpartition. Once both jobs start running, any subsequent job jd01 submits to that partition will be queued (i.e. placed inPENDING, orPD, status). Here is an example of how the output of thesqueuecommand would look like:JOBID PARTITION NAME USER ST TIME NODES NODELIST(REASON) 799 cpu384g A jd01 R 1-21:32:01 4 cpusm[01-04] 800 cpu384g B jd01 R 1-21:32:22 4 cpusm[05-08] 821 cpu384g C jd01 PD 0:00 4 (QOSMaxNodePerUserLimit)